Aimee Horsfall
Title: “DNP‐Supported Solid‐State NMR Spectroscopy of Proteins Inside Mammalian Cells”
Authors: Siddarth Narasimhan, Stephan Scherpe, Alessandra Lucini Paioni, Johan van der Zwan, Dr. Gert E. Folkers, Prof. Dr. Huib Ovaa, Prof. Dr. Marc Baldus
Journal: Angewandte Chemie International Edition
Year: 2019
https://onlinelibrary.wiley.com/doi/full/10.1002/anie.201903246
Featured image and figures used with permission under the Creative Commons Attribution‐NonCommercial‐NoDerivs License
Humans, chimps, zebras, rhinos, eagles, insects and sharks: All these different animals work together in a huge ecosystem, performing different jobs but each has their own role to make sure everything functions smoothly. Similarly, each cell in your body has its own thriving ecosystem of proteins, each with their own role to keep you functioning smoothly. Unlike studying the animal kingdom, where you can grab your camera and camp out behind a shrub, studying the proteins in your cells is far more difficult because they’re so small. Lucky for us, a bunch of really clever scientists have come up with a number of ways to help us understand cell biology. However, most of the methods used to look at proteins in the cell don’t show a lot of (molecular level) detail and obtaining details, such as the shape of a proteins, requires taking them out of the cell.
It is the special characteristics of each animal that gives rise to their special talents – a chimp swings from branches, a shark can swim rapidly through the water, or the eagle can fly long distances. Each of these talents is because of different ‘structural characteristics’ of each animal – the long arms of the chimp, the fins and gills of a shark or the wings of an eagle; and it’s the same for proteins – the 3D shape of a protein defines its function and its role in the cell. This is why it’s really important to be able to understand the shape of proteins, and more important to understand in the context of its environment – a jellyfish would look pretty silly out of the water. Until recently it’s not been possible to look at the structure of a protein while it’s still in the cell. Now we have techniques which are making this possible but it is still challenging to get good data at a concentration that is biologically relevant. This paper outlines a technique which can help enhance the signal, even at low concentrations, to achieve 3D structural data on a protein within a human cell.
Cells are really crowded, there’s a lot of different proteins and other ‘stuff’ going on in there. The first job is to tag the protein you are interested in looking at, this paper looked at a protein called ubiquitin which is a really important regulator protein. Tagging proteins allows us to observe only the protein of interest and tune out the rest of the crowd. Here, instead of using ordinary carbon and nitrogen, the isotopes C13 and N15 were incorporated into the protein structure. These isotopes, unlike their more naturally abundant partners C12 and N14 are “spin-active”, which means we can “see” or detect them using a technique called NMR.

The cells are then doped with this labelled protein through electroporation. This is where a small electric field is applied to the cells which makes the cell membrane more porous and allows the labelled protein to enter. Then the magic sauce is added, a DNP agent. This is a chemical which has a free radical, this is a spare electron, usually electrons hang out in pairs, but a radical is a lone electron. The electron can be transferred to the surrounding environment, and hopefully to the protein of interest – this becomes important in the next step. The sample of cells, doped with the labelled protein ubiquitin, is then snap frozen in liquid nitrogen and then subjected to a technique called solid-state Nuclear Magnetic Resonance Spectroscopy (ssNMR). This involves applying a large magnetic field applied to the sample, which is then pulsed with a radio-frequency wave that results in a change or a polarisation in the nucleus of each atom in the sample. As each atom relaxes back to its normal state of being, it releases a bit of energy that we can ‘collect’ with a detector – these pieces of information can be pieced together into a spectrum that tells us about how all those atoms are connected to one another and how they’re arranged in 3D space. As we’re only able to detect “spin active” isotopes, like C13 and N15, the majority of the signal we get from the sample should be from our labelled protein which is enriched with these isotopes – this lets us tune out the rest crowd. Solution-state NMR is very routine for chemists and used in almost every University chemistry department but solid-state NMR, like on this frozen sample, is more specialised.
Standard NMR samples are usually quite concentrated, e.g. 10-20 mg of sample per mL of solution. However, the frozen cells samples used by these researchers only have 10-30 ug of labelled ubiquitin – that’s 1000x less than normal and results in a very weak signal. This is where the magic sauce, the DNP agent, becomes important. The DNP agent allows for a technique called Dynamic Nuclear Polarisation to be performed. When the free radical is passed on to the labelled ubiquitin it causes a more severe polarisation of the atomic nuclei than the NMR is able to achieve alone. The knock-on effect of this is that the signal is enhanced by more than 100x. Though this technique has been used before, these researchers are the first to demonstrate it to study protein structure inside a human cell. The DNP agent is not very stable, and this is made worse by the reductive environment inside a human cell. However, snap-freezing and conducting the experiment in the solid-state, rather than the solution state, allowed these researchers to overcome this problem!
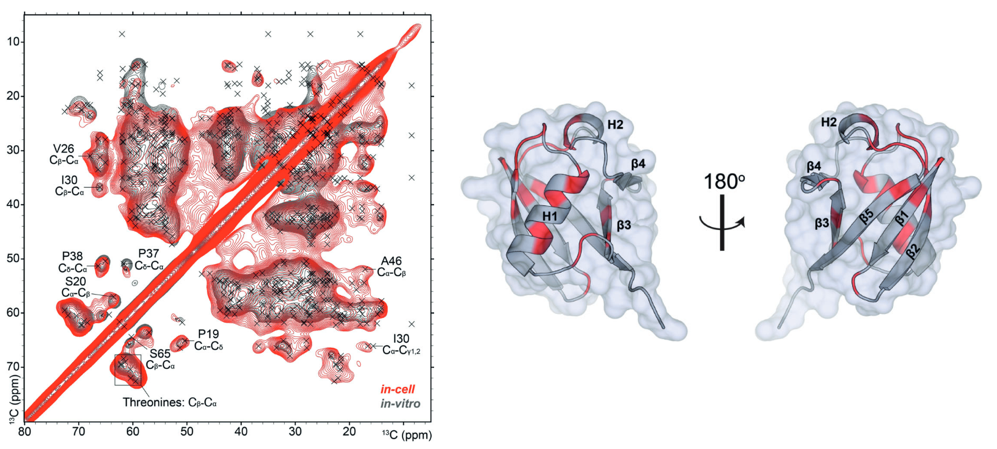
This is a really exciting technique which provides researchers another set of tools to study proteins in their natural environment, and allow us to see whether these proteins take on different conformations while interacting with components of the cell, whether a protein is malformed and causing a particular disease, or even how different signalling agents or drugs interact with important proteins. Hopefully this will allow us to better understand diseases, identify new drug targets and more rapidly design and test therapeutics.

