Title: Computational Prediction of Electron Ionization Mass Spectra to Assist in GC/MS Compound Identification
Authors: Felicity Allen, Allison Pon, Russ Greiner, David Wishart
Published: 6 July 2016
Journal: Analytical Chemistry
http://pubs.acs.org/doi/abs/10.1021%2Facs.analchem.6b01622
If you want to know about a molecule without chemicals, now you can without entering a lab.
A common practice in knowing a molecule is like solving a puzzle pieces by pieces. Therefore, the first thing to do is to shatter the molecule into pieces, then analyse the fragments to get the whole picture. The abundance and mass of the fragments produce a spectrum that becomes the fingerprint of the molecule. This is often done by electron-ionisation mass spectrometry (EI-MS).
Now, these can be achieved by computational power.
Predicting a molecule with algorithms has been a dream of many. However, the path to realize this dream is full of thorns. One way is to identify the possible pieces of the target molecule fragments. This can be done by setting various constraints in algorithms to generate likely structural isomers of the target. However, this could result too many candidate structures of the target than computational power can manage. A way to tackle this problem is by setting up more rules in the calculations, but the difficulties lay in the choice of rules.
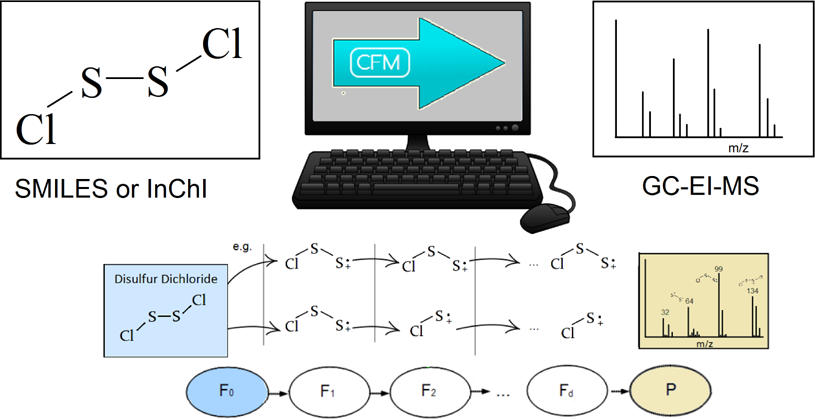
Instead of setting more rigid rules, the authors here introduce another approach, which involves breaking every bond of the target molecule systematically and recursively. They called it the called competitive fragmentation modelling (CFM). In the example of a disulfur dichloride molecule, every pair of bonds is broken and charges and radicals are rearranged within each pair of resulting fragments (Figure 1). The transition function in the algorithm describes the neural network of sequence of breaking bonds and the observation function is for handling isotopes. Another feature of their algorithm is taking into account of odd-electron ions as the target molecule in EI-MS contains odd number of electrons.

Figure 1 The schematic diagram of the CFM-EI procedure.
This CFM-EI model is one of the few methods to predict the electron-ionisation spectrum of the molecule with mass-to-charge ratio. They have verified this method against NIST database where their correct candidate scored rank 1 in 77% of cases. Their model was also successful in many metabolite identification tasks, demonstrating its capability in small molecule prediction.
Though there may still be a chasm between the computationally predicted and real measured spectra, computational method is a rapid and cost-effective alternative for collecting reference spectra of chemical standards. Since science is a collaborative effort, the authors have generously shared their source code at https://sourceforge.net/projects/cfm-id for those having interests.

Pingback:Get to know about molecules without chemicals! – Eli's Science Blog