Scientific Article Information
Title: Identification of a S. aureus virulence factor by activity-based protein profiling (ABPP)
Authors: Christian S. Lentz, Jessica R. Sheldon, Lisa A. Crawford, Rachel Cooper, Megan Garland, Manuel R. Amieva, Eranthie Weerapana, Eric P. Skaar & Matthew Bogyo
Journal: Nature Chemical Biology
Year: 2018
https://www.nature.com/articles/s41589-018-0060-1
Staphylococcus aureus is a beneficial bacterium living on our skin and mucus. However, when S. aureus ends up in the wrong place, it can become a pathogen leading to life-threatening conditions like meningitis and sepsis. A key aspect of its pathogenicity is the ability to form biofilms, which are communities of microorganisms that adhere to a surface. S. aureus biofilms serve as a wall that blocks antibiotics and our immune system. A better understanding of the factors that contribute to S. aureus biofilms may inform improved treatment options.
Serine hydrolases are a class of enzyme that play important roles in host-pathogen interactions, such as those that occur when S. aureus forms biofilms and evades the human immune response. Importantly, scientists have a long history of designing drugs that block this class of enzymes. However, the functions of serine hydrolases in S. aureus were not well-characterized. In this article, Lentz et al. used a method called activity-based protein profiling to identify serine hydrolases important for S. aureus virulence.
Activity-based protein profiling
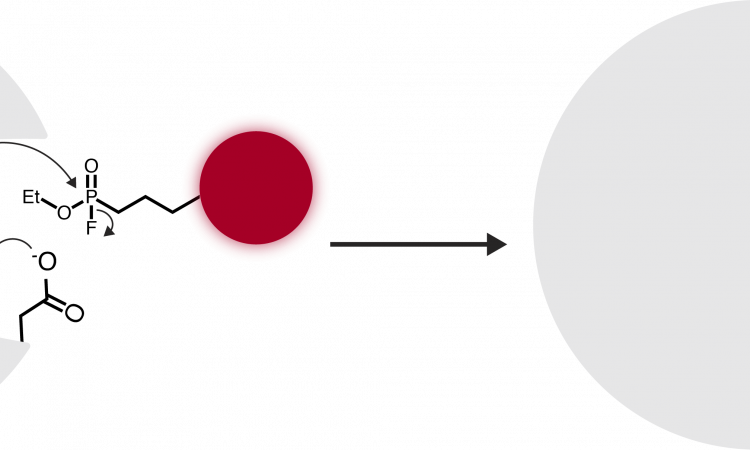
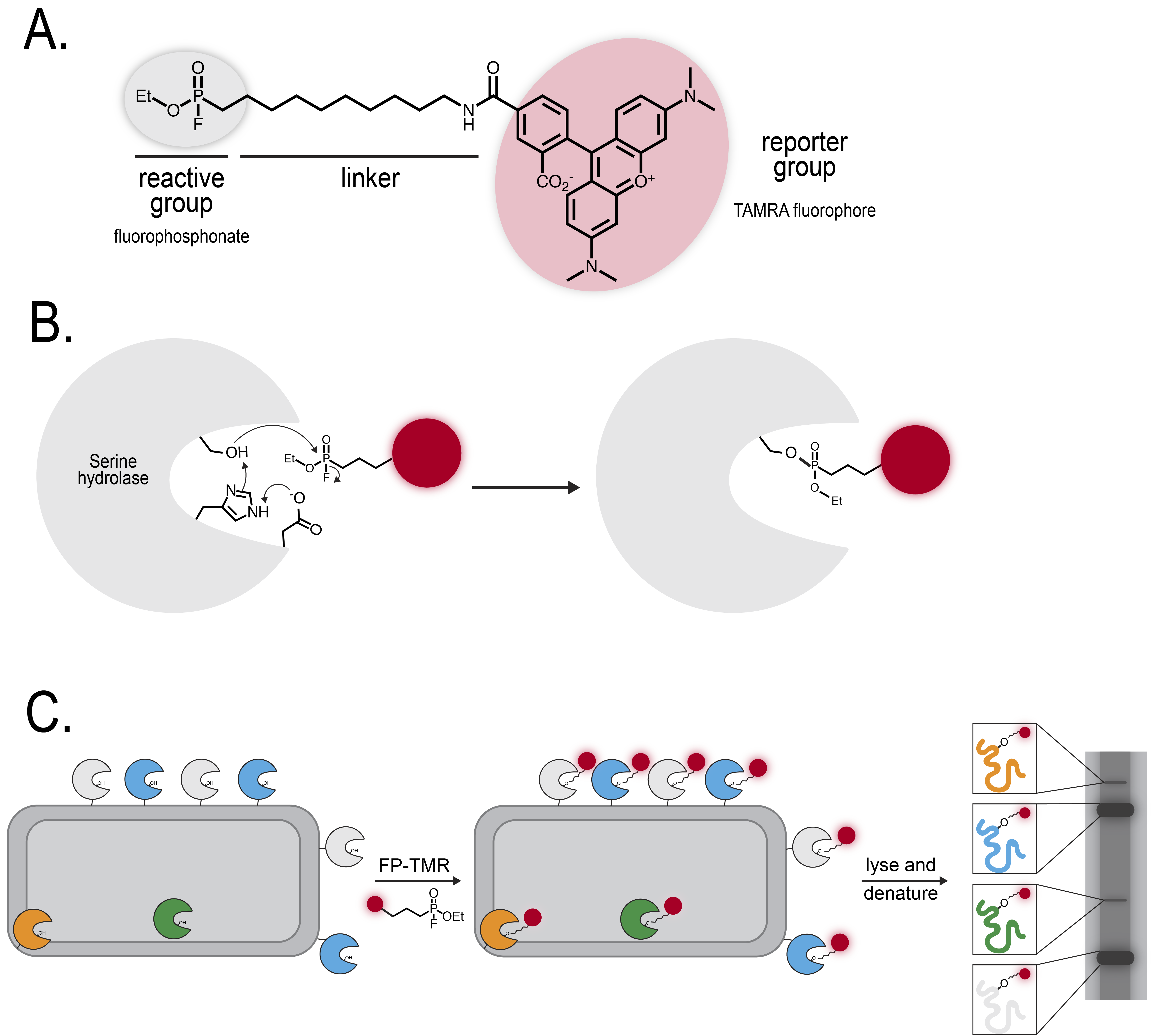
Activity-based protein profiling (ABPP) capitalizes upon enzyme mechanisms involving covalent intermediates, such as the acyl intermediate formed by serine hydrolases. The chemical ABPP probe consists of reactive, linker, and reporter groups (Figure 1A). First, the enzyme must be active to bind the probe. Next, the probe functions as a “suicide inhibitor”, trapping the enzyme in the intermediate state (Figure 1B).
Fluorophosphonates have commonly been used as activity-based probes for serine hydrolases. This study used Fluorophosphonate tetramethylrhodamine (FP-TMR), which can be visualized on a gel (Figure 1C). First, the researchers performed a “functional chemoproteomic screen”: they treated bacteria isolated from biofilm-promoting conditions with FP-TMR and visualized labeled proteins on a denaturing gel. This experiment revealed many candidate serine hydrolases that may play a role in maintaining biofilm conditions. To dig deeper into the role of specific serine hydrolases, however, the researchers had to perform a competitive ABPP experiment.
Using competitive ABPP to identify specific S. aureus serine hydrolases
A competitive ABPP experiment allows researchers to identify specific inhibitors of an enzyme. In this case, a specific inhibitor refers to a small molecule that predominantly inhibits only one serine hydrolase. Cells are first pre-treated with a chemical inhibitor and then treated with FP-TMR (Figure 2A). If the inhibitor binds a serine hydrolase, then the inhibitor will prevent the FP-TMR from binding the same serine hydrolase. As a result, the fluorescence intensity of the gel band corresponding to that serine hydrolase will decrease as the concentration of specific inhibitor increases.
The researchers performed competitive ABPP experiments using a repository of ~500 compounds. They identified two compounds that appeared to inhibit specific hydrolases. To identify the protein “targets” inhibited by these chemicals, the researchers switched their probe from FP-TMR to FP-biotin (Figure 2B). Biotin is a small biological compound that is tightly bound by certain types of proteins like streptavidin. By attaching streptavidin to beads, scientists are able to “pull down”, or isolate, biotin – which, in this case, is covalently attached to a serine hydrolase (Figure 2C). Once the researchers isolated the serine hydrolases, they could assign their identities using a method called mass spectrometry.
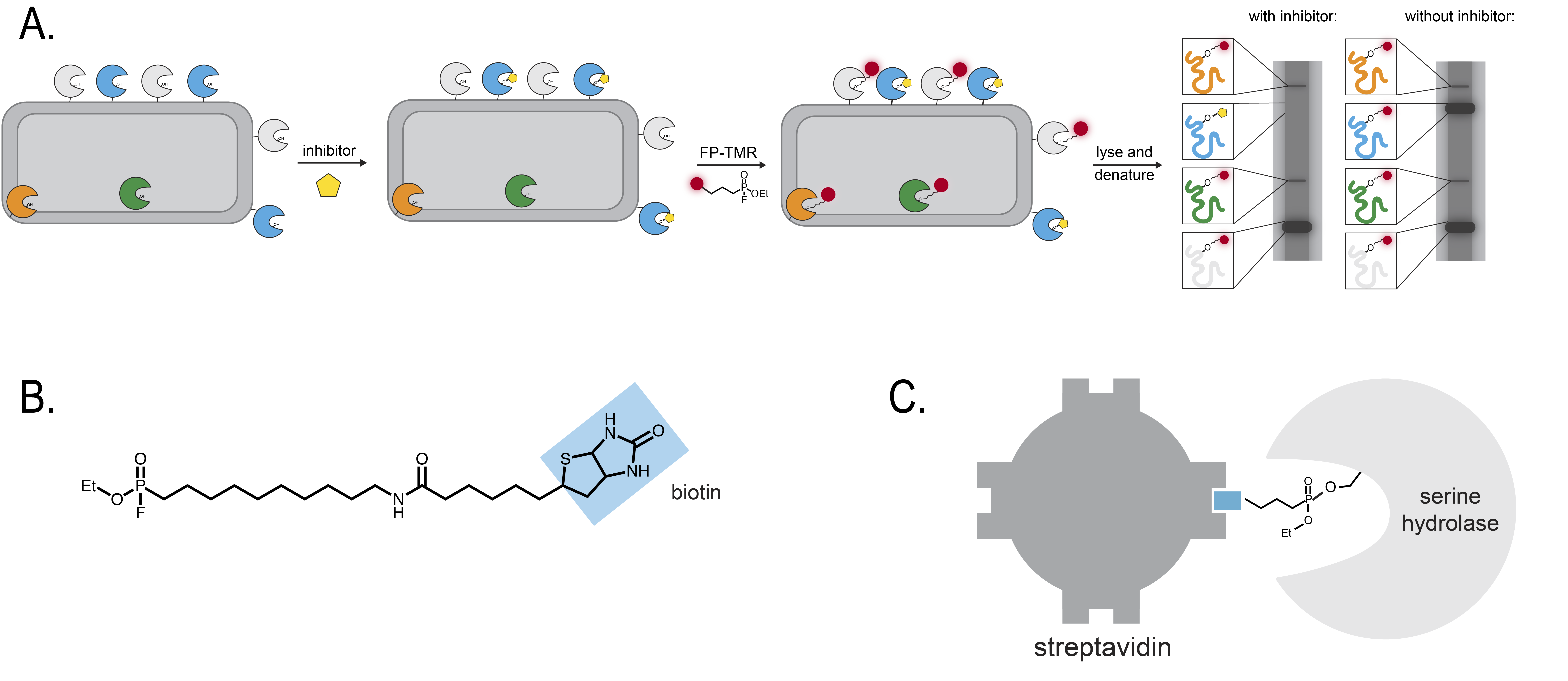
Figure 2. Experiments to identify specific serine hydrolases. A. In a competitive ABPP experiment, cells are first incubated with an inhibitor and later with the ABPP probe, in this case FP-TMR. If the inhibitor binds a specific enzyme, then the enzyme will not react with the ABPP probe. As a result, the band on the fluorescence gel becomes less intense. B. Biotin is a small molecule that can serve as a reporter group of an ABPP probe. C. Streptavidin strongly binds biotin. If a serine hydrolase reacts with a biotin ABPP probe, then purifying streptavidin bound to biotin n will also purify the serine hydrolase.
Using this approach, the researchers identified 10 proteins, which they called “fluorophosphonate-binding serine hydrolases”, or Fphs. Many of these proteins were “unannotated”, which means their cellular functions are unknown. The quest, therefore, is to figure out what these proteins do.
Biochemistry and chemical biology shed light on the role of a specific serine hydrolase
To verify that the candidate protein was a specific target of the fluorophosphonate probe, the researchers ordered mutant bacteria lacking functional hydrolases. When the profiling experiment was repeated on the mutant bacteria, the bands corresponding to specific hydrolases were missing, which showed the researchers that their candidate proteins were true targets of the probe.
The researchers then focused on figuring out where a specific hydrolase, FphsB, was localized and what it was doing. The researchers localized the hydrolase by creating a new, more specific fluorescent probe called JCP251-bT. When the bacteria were treated with JCP251-bT, some of the cell rims fluoresced, suggesting that FphsB is at the cell envelope. The researchers then dissected the function of the hydrolase by purifying the enzyme and incubating it with esterase, phosphate, phosphonate, glycosidic, or protease substrates. The purified hydrolase only cleaved the esterase substrates, suggesting that FphB may cleave fatty acids.
Conclusions and Next Steps
This study demonstrates the strengths of the ABPP approach. Starting with little knowledge of the identity or function of serine hydrolases in S. aureus, the researchers were able to identify hydrolases that appear to play a role at the host-pathogen interface and can be specifically inhibited by small molecules. The researchers then focused on a specific serine hydrolase, FphB, and identified its potential location and substrates. There is still a lot of work to be done: the true, physiological substrates of FphB are not known, which greatly limits our knowledge of the role it may play during infections. Furthermore, there are many more Fphs waiting to be characterized. Nonetheless, this approach, and ones like it, may be used to investigate many more previously unexplored classes of enzymes in the future.