Article Title: Determinants for substrate recognition in the glycyl radical enzyme benzylsuccinate synthase revealed by targeted mutagenesis
Authors: Salii, I.; Szaleniec, M.; Zein, A. A. et al.
Journal: ACS Catal.
Year: 2021
DOI: 10.1021/acscatal.0c04954
With increasing global dependence on oil, more and more oil spills occur throughout the world. These disastrous events pollute the air, land, and water with toxic hydrocarbons (chemicals composed of only carbon and hydrogen atoms) such as benzene and toluene. Microbial or enzymatic bioremediation efforts, or the use of microbes or enzymes to treat polluted areas, could be the solution. Bacteria such as Thauera aromatica K172 and Aromatoleum toluolicum T are known to anaerobically degrade toluene. They encode the enzyme benzylsuccinate synthase (BSS), which activates toluene and adds it to the co-substrate fumarate, forming benzylsuccinate (Figure 1A).
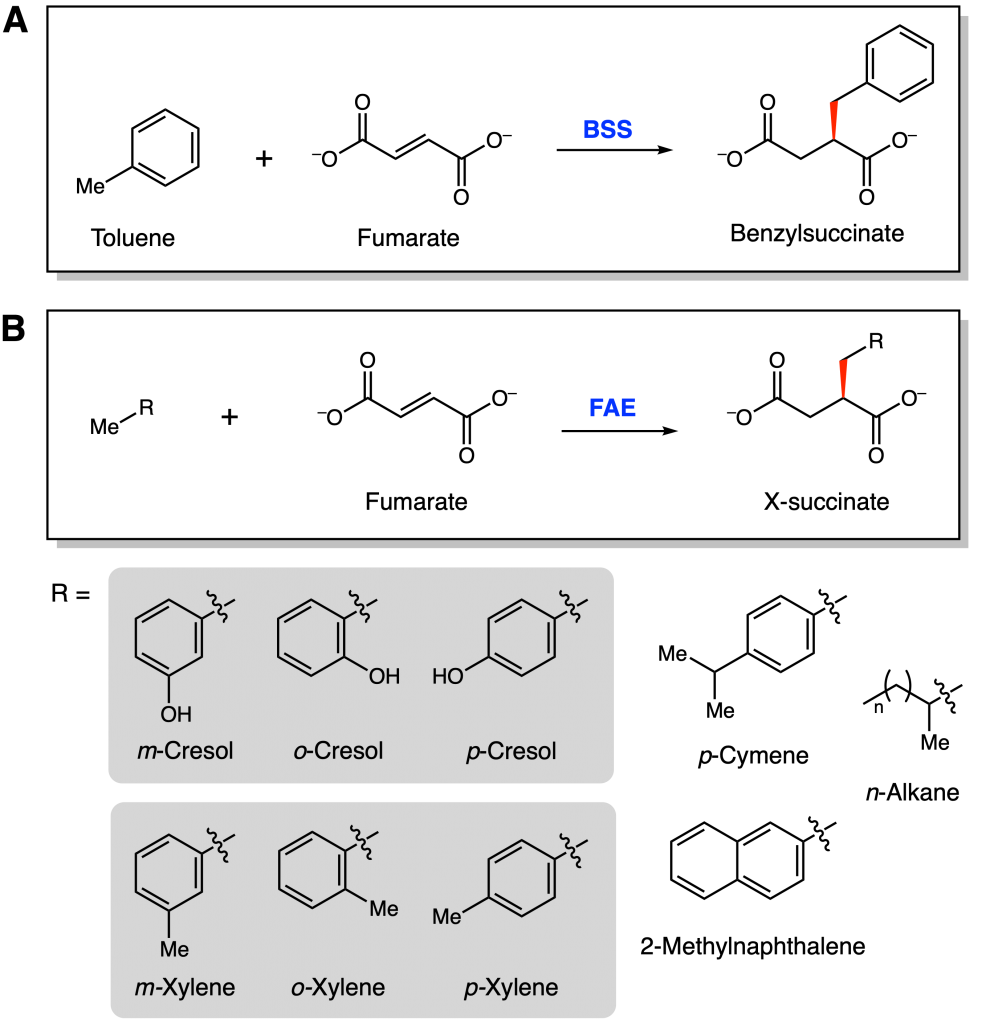
BSS is part of a larger fumarate-adding enzyme (FAE) subfamily whose members can also degrade various cresol, xylene, cymene, naphthalene, and alkane hydrocarbons (Figure 1B). All of these enzymes use radical chemistry, or 1-electron chemistry, to catalyze these difficult transformations, making them particularly useful in anaerobic environments such as deep soil. The FAEs have limited substrate scopes, meaning they only accept a few types of substrates. For instance, wild-type, or the natural form, T. aromatica BSS can metabolize toluene and cresols, but it cannot process xylenes.
In this paper, Salii and coworkers used targeted mutagenesis (or the changing of specific amino acids within a protein sequence) to alter the substrate scope of T. aromatica BSS. In particular, they targeted two isoleucine (Ile) residues that border toluene and an arginine (Arg) residue proposed to bind fumarate. The specific mutations are shown in Figure 2.
Before they could test the BSS mutants, they had to first improve the heterologous expression of BSS in a non-native bacteria. Previous attempts to use Escherichia coli as an expression host resulted in only inactive protein. Instead, the group used the bacteria Aromatoleum evansii, which is more closely related to the natural BSS producing organisms. Only cell extracts of A. evansii expressing BSS, and not A. evansii itself, were able to degrade toluene. Using this platform, different point mutants and their substrate scope could be tested and compared to the wild-type BSS.
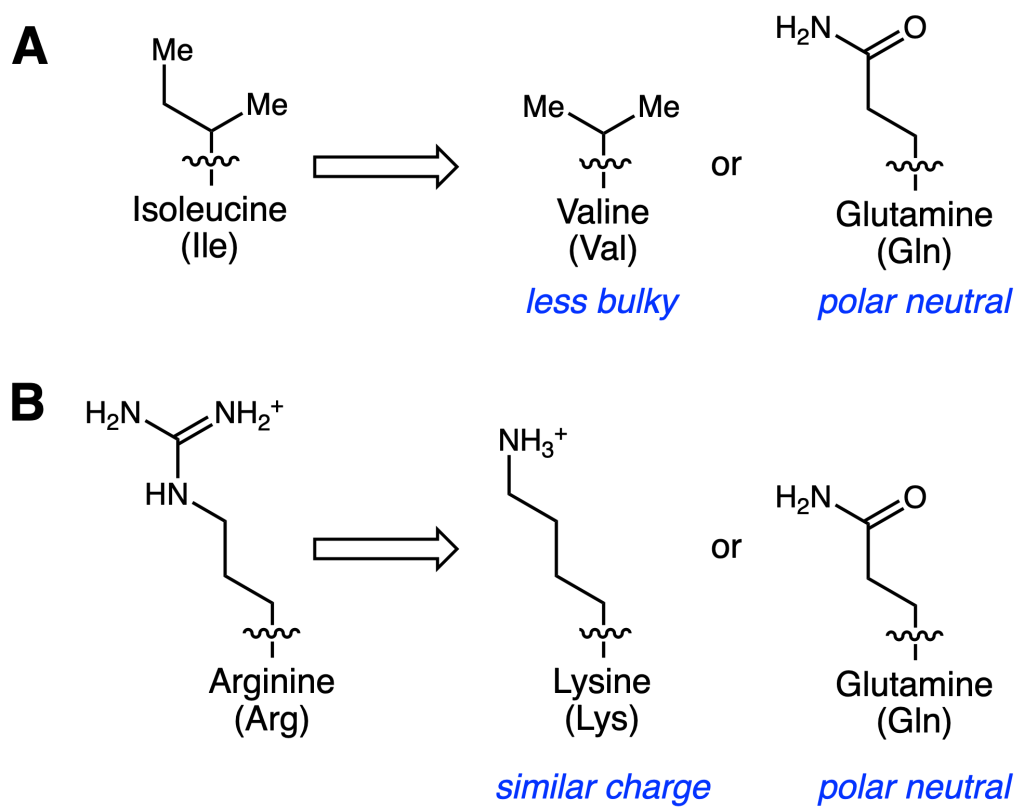
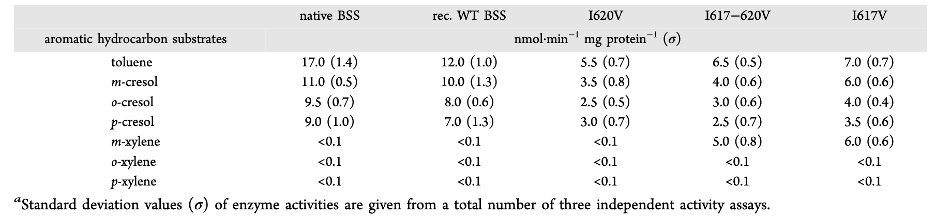
First, they mutated Ile at position 620 (Ile620) into glutamine (Gln), a larger, polar residue. Unsurprisingly, this large polarity perturbation resulted in inactive BSS. Mutants of Ile617 and/or Ile620 into the less bulky residue valine (Val), however, still were active. Toluene as well as a panel of cresols and xylenes were tested, and their specific activities are reported in Table 1. The specific activities for wild type toluene and the cresols decreased for all mutants. However, variants with the Ile617 to Val mutation are now able to metabolize m-xylene.

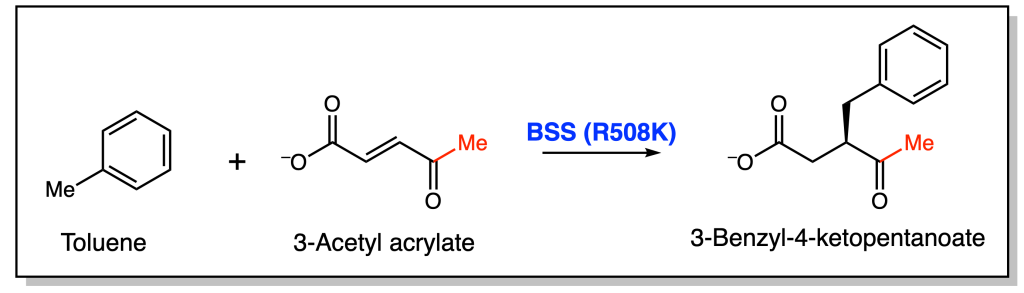
The other position they investigated was Arg508. The positively charged Arg is predicted to form a salt bridge with the negatively charged fumarate. As a result, they tested another positively charged residue, lysine (Lys), as well as the neutral, polar residue Gln. Only the Lys variant was active towards toluene, albeit with lower specific activity. This suggests that the proposed salt bridge is important for BSS’ activity. Surprisingly, when the Lys variant was incubated with 3-acetyl acrylate instead of fumarate, small amounts of product appeared to form (Figure 3). To corroborate these findings, the authors used molecular dynamics (MD) simulations to compute how different BSS mutants bind the alternative substrates.

In the end, the authors were able to show that rational mutagenesis of BSS can expand its substrate scope to degrade a variety of hydrocarbons. Although few positions were tested, one of the mutations, I617V, enabled BSS to process m-xylene with similar efficiencies as the native toluene substrate. It will be exciting to see if further work exploring a broader range of mutations throughout the ~860 long protein sequence will lead to more efficient enzymes with enhanced substrate scope.

