Article Title: Fluorinated S-adenosylmethionine as a reagent for enzyme-catalyzed fluoromethylation
Authors: Peng, J.; Liao, C.; Bauer, C.; Seebeck, F. P.
Journal: Angew. Chem. Int. Ed.
Year: 2021
DOI: /10.1002/anie.202108802
Of the many elements in the periodic table, fluorine is one of the most privileged and sought-after functional group to introduce into drugs. Although fluorine is similar to hydrogen, the replacement of hydrogen with fluorine can dramatically alter drug efficacies by changing its solubility and stability within cells. Due to these desired principles, many lab groups have been developing both synthetic and biological means to introduce fluorine into various compounds. Here, Peng and Liao et al. report using the enzyme halide methyltransferase (HMT) as a way of incorporating the functional group monofluoromethyl, an analog of the methyl group, into various small molecules (Figure 1A).
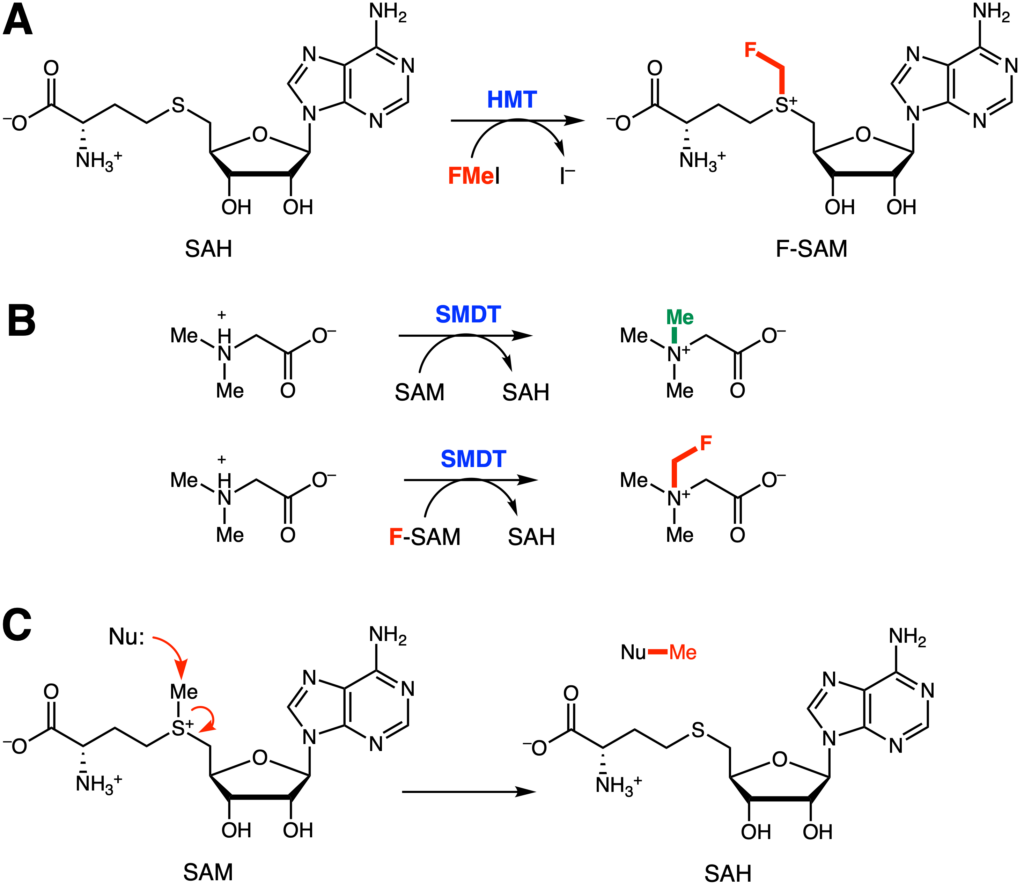
The key step is generation of a fluorinated cofactor, F-SAM, that can be used by other enzymes to attach fluoromethyl rather than the usual methyl groups (Figure 1B). S-adenosylmethionine (SAM) is a common cofactor involved in enzymatic methylation reactions (Figure 1C). In a typical reaction mechanism, a nucleophile will attack the terminal methyl group, pushing electrons onto the electron-deficient sulfur and generating the byproduct SAH. Halide methyl transferases (HMTs) are a common class of enzymes used to create SAM analogs. For instance, they have been engineered to transfer ethyl, propyl, allyl, and carboxylated alkyl groups. The authors wanted to expand the enzyme’s chemical potential and thus investigated whether fluoromethyl groups could also be appended.
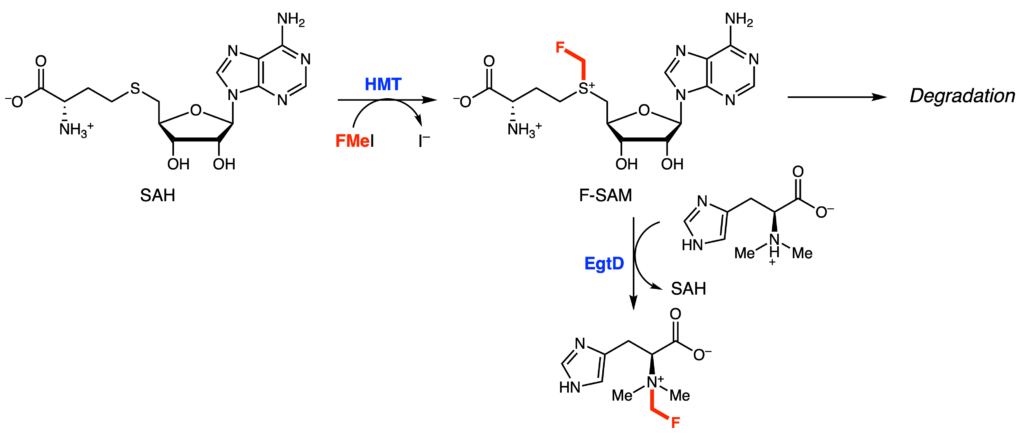
Specifically, they worked with a Burkholderia xenovorans HMT, which upon incubation with fluoromethyl iodide (FMeI) and SAH, led to the production of small amounts of F-SAM. However, the majority of compounds produced were actually decomposition products of F-SAM, as it is easily hydrolyzed. Given the fleeting nature of F-SAM, the authors wondered if in situ generation of F-SAM could be used by a different methyltransferase faster than the rate of degradation. In other words, if HMT and a second methyltransferase were mixed together in one pot, could the small amounts of F-SAM produced in the reaction serve as a cofactor for the second methyltransferase. To test this, they added the methyltransferase EgtD and its substrate, dimethylhistidine. Satisfyingly, they saw formation of F-trimethylhistidine (Figure 2), implying that EgtD successfully outcompetes the rate of F-SAM degradation.
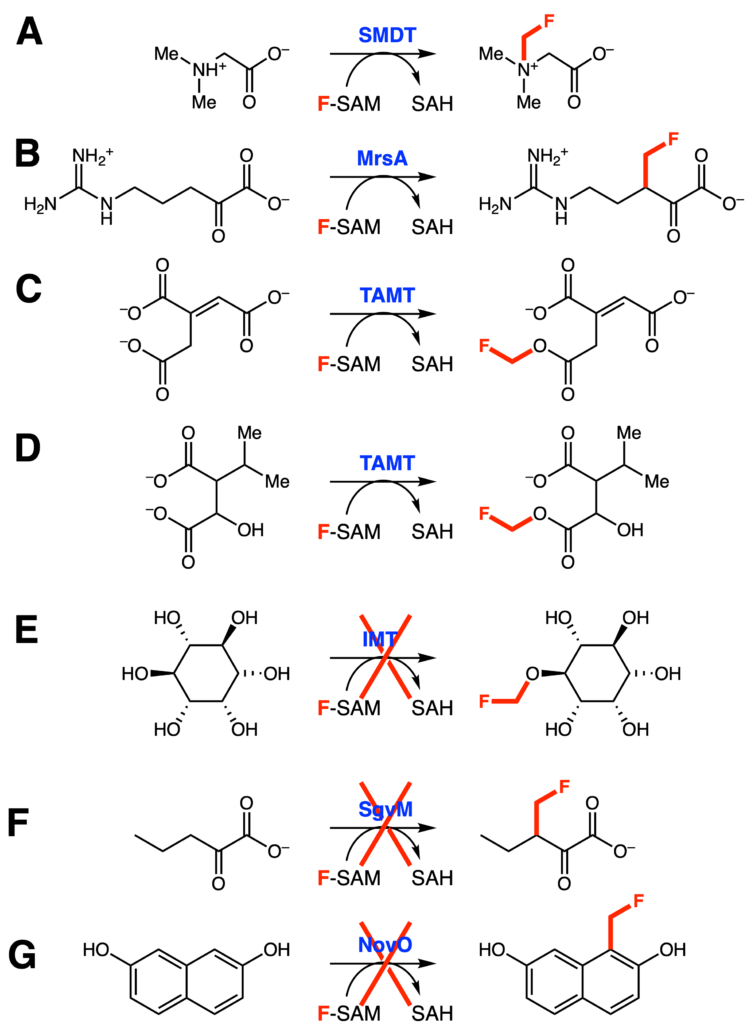
This same workflow was then expanded upon to test a variety of unrelated methyltransferases (Figure 3A-F). Five of eight enzymes tested (including EgtD in Figure 2) were able to use F-SAM as a fluoromethyl donor. Multiple nucleophile identities (carbon, nitrogen, and oxygen) were all able to append a fluromethyl group. The reactivity, however, is not universal, as no fluorinated products of IMT, SgvM, or NovO were detected. The authors postulated that this is due to differences in reactivity rather than size preferences. Both fluorine and hydrogen are similar in atomic radius, and SgvM and NovO both can utilize the SAM analog that has the bulkier ethyl group in place of the methyl group.
In this work, an enzymatic method of installing fluoromethyl groups was developed. This critically relies on generation of transient amounts of F-SAM by HMT that is then quickly consumed by downstream methyltransferase enzymes. The small subset of methyltransferases tested demonstrates clearly that many enzymes are innately promiscuous and can transfer fluromethyl groups to their native substrate. Besides the methyl functional group, fluoromethyl is similar in size and shape to primary alcohols, thiols, and terminal amines. It will be exciting to see how these fluorinated small molecule analogs alter the function of the small molecules, as many of them have antibiotic or therapeutic potential.