Title: A Naturally Occurring Antiviral Ribonucleotide Encoded by the Human Genome
Authors: Steven C. Almo et al.
Journal: Nature
https://www.nature.com/articles/s41586-018-0238-4
Year: 2018
Viral infections continue to challenge healthcare and wellbeing around the globe. In the ever-demanding game of evolution, a host needs to develop sophisticated biological machinery that will target an essential process in the virus, and will often have to respond swiftly to the virus’ evolutionary comeback. Inherently, Nature follows the same principles when adapting to viral infections. One of such weapons we deploy to combat viral infections is a protein known as viperin. It has been shown in the past that viperin can inhibit the replication of many RNA- and DNA-based viruses, yet the exact means by which it does so remains obscure… until now. In this paper, a collaboration between research groups at the Albert Einstein College of Medicine and Penn State University finally determine the unique mechanism by which viperin inhibits viral replication.
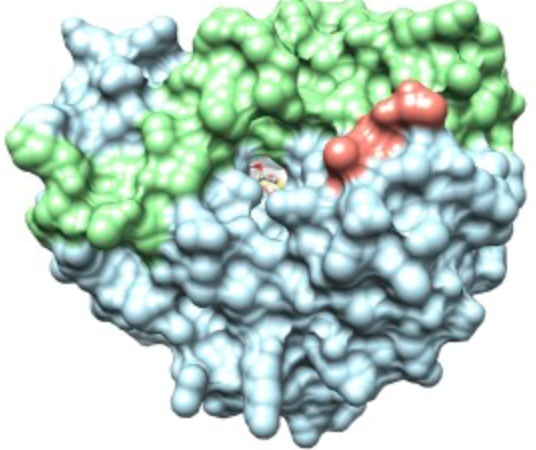
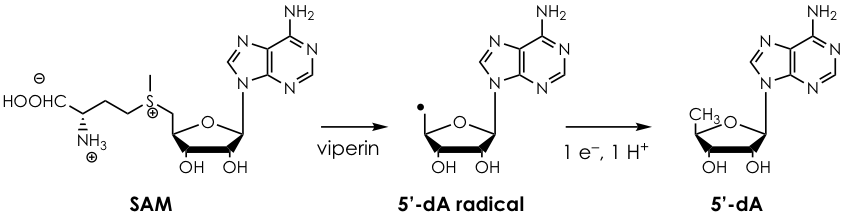
The authors first observed that (i) the gene coding for viperin is almost always present next to a gene coding for a protein that modifies cytosine and (ii) that viperin is an enzyme which used a cofactor known as S-adenosylmethionine or SAM (Figure 1). This suggested that viperin is most likely a nucleoside-modifying enzyme and that it operates via radical SAM chemistry (Figure 1). To initially test their hypothesis, they first screened the purified protein against a collection of nucleotides and checked for enzymatic activity by looking for one of the products of SAM chemistry, 5-deoxyadenosine (5’-dA). They observed that only cytosine triphosphate (CTP) activates viperin to a reasonable extent and that the reaction generates another product.
Further analysis using mass spectrometry and UV-Vis and NMR spectroscopy revealed this product to be 3’-deoxy-3’,4’-didehydro-CTP or ddhCTP (1, Figure 2). To define the mechanism by which 1 is formed, the authors incubated the enzyme with a synthetic derivative of CTP that has deuterium atoms (heavier versions of hydrogen) incorporated at key places. Then, using very sensitive mass spectrometry techniques, they can piece together the chemical steps that take place in this transformation (Figure 2).
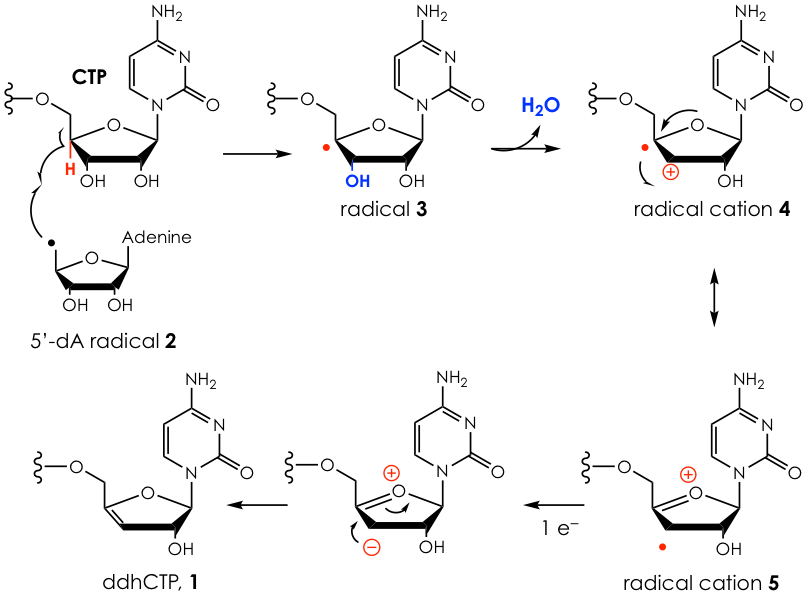
In short, the enzyme first generates 5’-dA radical (2) from SAM. This radical then selectively abstracts the hydrogen at the 4’ position of the CTP ribose to get radical 3. The 3’ hydroxyl is then removed with the help of a general acid to produce the radical cation 4, which can also be resonance-stabilized to 5. This intermediate then receives one electron and rearranges to yield the product 1. It is unclear still where that extra electron comes from.
Next, the authors validated the presence of 1 in mammalian cells and used the Dengue virus as a model to show that 1 acts as a terminator of viral RNA polymerase. Viruses use this enzyme to stitch together ribonucleotides into long RNA strands, but when 1 instead of CTP is incorporated, the polymerase cannot further elongate the strand, and the virus cannot replicate accurately! The curious reader is encouraged to visit the paper to get details on the assays and results leading to this exciting discovery.
There are still many questions that remain unanswered. The chemical mechanism for the formation of 1 proposed here, for instance, is only partially validated, and it remains to be seen if the biological mode of action of viperin is conserved against other viruses. Still, these findings are novel and impactful and will probably represent a significant milestone in the field of viral resistance.