Title: Monitoring the Immunoproteasome in Living Cells Using Activity-based Peptode-peptoid Hybrid Probe
Authors: Darci J. Trader and Breanna L. Zefas
Journal: JACS
Year: 2019
https://pubs.acs.org/doi/10.1021/jacs.8b12873
Cells house an impressive collection of intricate macromolecular structures with dedicated roles in maintaining homeostasis. The ribosome, for instance, is a mega-Dalton assembly of RNA and protein in charge of directing protein synthesis, and the so-called p-bodies are foci of many enzymes critical for the decay of unused or defective messenger RNA. Another one of these structures – the proteasome – is a multiprotein complex is in charge of hydrolyzing (degrading) other proteins no longer needed by the cell, thus taking an elemental role in cellular protein level balance.
In immune cells, a slightly different version of the standard proteasome – the aptly named immunoproteasome – is expressed when these cells encounter inflammatory elements and is known to be overexpressed in certain cancers and autoimmune disorders (Figure 1). There exist very few tools to study the levels and activity of this structure in live cells, let alone tools that selectively interrogate the immunoproteasome over its standard cousin.

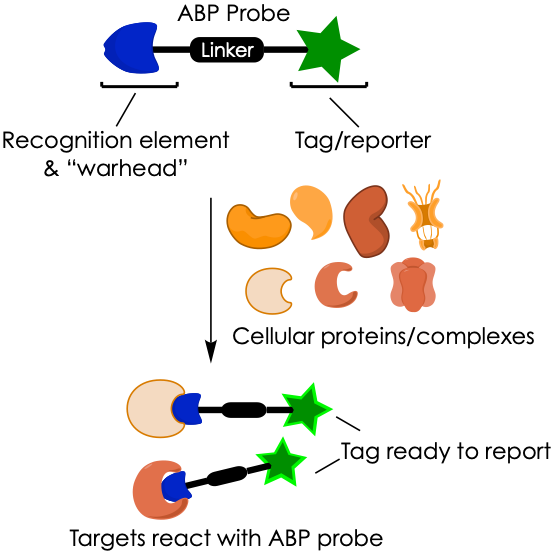
To address this shortcoming, the group of Darci Trader at Purdue University recurred to activity-based protein profiling (ABPP). This approach has become a mainstream technology in chemical biology and consists of using a small-molecule probe that can be taken up by cells and selectively report on the activity of a specific enzyme or macrostructure. To do so, the ABP probe must contain (a) a directing element that mediates interaction with the cellular target, (b) a reactive moiety (also known as “warhead”) that engages with the target in a way similar to how the target would react with its native substrate, and (c) a tag that reports once reaction with the target (i.e., activity) has taken place (Figure 2).
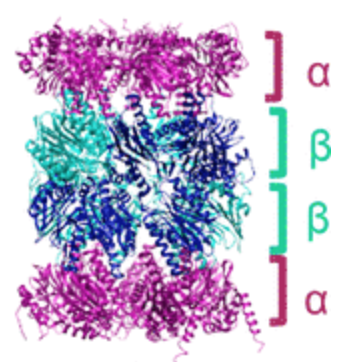
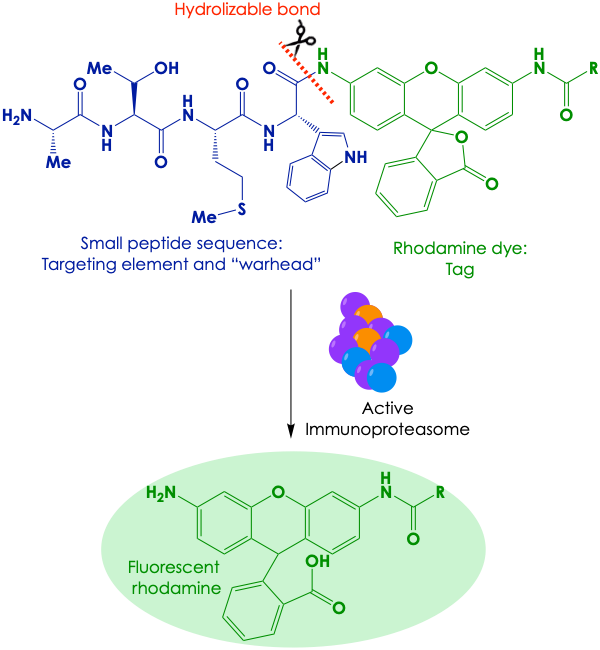
To aim at the immunoproteasome, the authors first focused on previously solved crystal structures of this complex and of its conventional counterpart. They looked for differences in the catalytic subunit that could aid in the design of immunoproteasome-selective directing elements of the ABP probe. After screening for immunoproteasome-mediated hydrolysis of a library of candidate probes, they settled with a tri-peptide motif with bulky and hydrogen-bonding amino acids. This motif binds to and is hydrolyzed by the immunoproteasome quite well, and so, it can function both as targeting element and “warhead” (Figure 3). For the reporter, the authors appended a rhodamine fluorophore at the hydrolyzable position. The intact probe is non-fluorescent, but when an active proteasome performs hydrolysis at the key bond, free rhodamine is released and readily isomerizes to its fluorescent form.
With the probes in hand, the authors first confirmed the probe’s fluorogenic reporting in vitro through a hydrolysis assay with the isolated immunoproteasome. They observed increased fluorescence for more prolonged reactions since more free rhodamine is generated over time.
In vivo, the team was able to show the effectiveness of their ABP probe. They cultured immune cells and treated some of them with an inflammatory agent known as interferon-gamma (INF-gamma). Next, they fed all cells with the ABP probe and observed the cells under a fluorescent microscope. As expected, fluorescence was always higher in cells treated with IFN-gamma given they overexpress immunoproteasomes.
While there are many details from the original paper left behind in this recount, it is not hard to appreciate the potential of this new ABP probe in fundamental chemical biology and even in the biomedical arena. The immunoproteasome could become a well sought-after target in cancers and autoimmune diseases, and only time will tell how enabling this tool will become then.