Title: Surveying the Vampire Bat (Desmodus rotundus) Serum Proteome: A Resource for Identifying Immunological Proteins and Detecting Pathogens
Authors: Benjamin A. Neely, Michael G. Janech, M. Brock Fenton, Nancy B. Simmons, Alison M. Bland, and Daniel J. Becker
Journal: ACS Journal of Proteome Research
Year: 2021
https://doi.org/10.1021/acs.jproteome.0c00995
Featured image adapted from Pixaby free-use images.
Spring 2021 has been a strange time, with many feeling relief as they receive the COVID-19 vaccine, while others around the world are still struggling through the midst of the pandemic. Still, the speed at which multiple vaccines were developed highlights the lifesaving power of scientific research. As many look hopefully toward the future, some scientists are developing methods to better track viruses in animals to better understand the causes of zoonotic transmission (the transmission of a disease from animals to humans). According to the CDC, 3 out of 4 emerging infectious human diseases originate in animals [1]. In the case of SARS-CoV-2 (the viruses that causes COVID-19), the World Health Organization (WHO) report showed that zoonotic transmission was likely, but WHO was unable to determine the exact origin of the disease [2]. Some evidence showed that animals sold at markets close to the original cases were farmed in regions where bats and pangolins carried viruses similar to SARS-CoV-2 [2]. However, none of these viruses were similar enough to be the direct source of the COVID-19 virus [2], and the true origin remains a mystery.
While bats may or may not carry the virus of this pandemic, they are known to carry many other zoonotic viruses including other SARS-like coronaviruses, herpesvirus, adenovirus, and the rabies virus. Currently, the country of Belize is particularly concerned with the increasing number of rabies outbreaks in livestock since 2016. Virus samples taken from livestock matched those seen in vampire bats which feed on blood from livestock, giving them an opportunity to transfer pathogens. To improve future efforts to detect viruses in bats, scientists Neely and coworkers decided to see if the increasingly popular “shotgun proteomics” technique would be useful in identifying all the proteins—and therefore all viral proteins—circulating in vampire bat blood. Here, the suffix “omics” refers to finding “all” proteins, and the term “shotgun” refers to the quick speed of the technique.
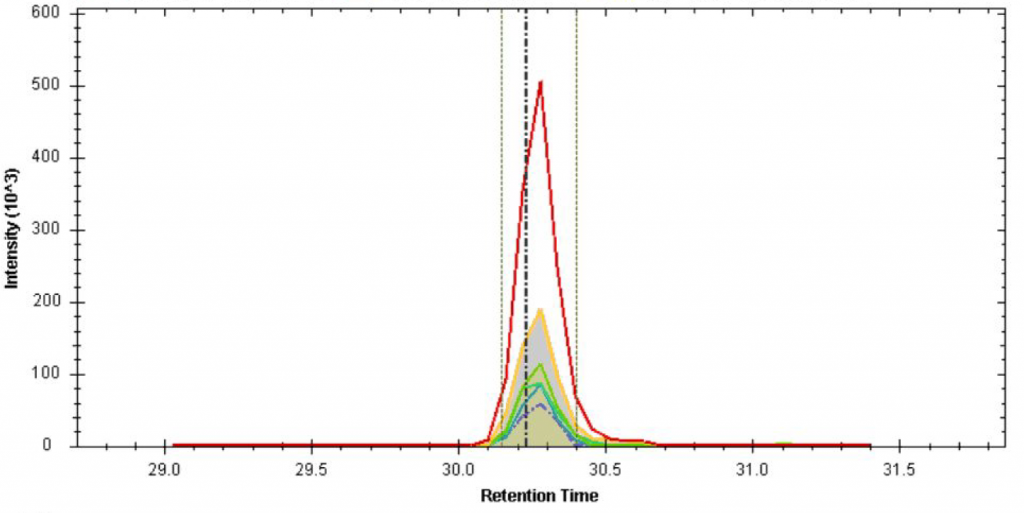
Different viruses use unique proteins to carry out their infections within hosts, such as the Spike protein in SARS-CoV-2 that allows the virus to gain entry to human cells. Neely and coworkers wanted to search all the proteins circulating in the bloodstreams of vampire bats to find viral proteins, thus identifying which viruses the bats were carrying. The scientists used blood samples from 17 bats in Belize that were already being tracked for a long-term study related to the rabies virus in the area. The blood was clotted to remove all the cells (e.g. white and red blood cells and platelets) from the samples, and the cell-containing clots were removed, leaving blood serum behind and ready for the next steps: protein digestion followed by Liquid Chromatography-Mass Spectrometry (LCMS, Fig. 1).
Because whole proteins are too big to be detected and analyzed easily, an enzyme called trypsin was used to break the proteins down into smaller pieces called peptides. Trypsin is an enzyme our own bodies use to digest protein after we eat, and it does the same job in the lab. At this point, there were too many peptides to all be detected at once, so they were separated by a method called liquid chromatography (LC). In LC, the peptide samples flowed through a column containing anchored hydrophobic molecules (similar to butter or oil). This separated different peptides from each other, similar to how oil separates from vinegar in a salad dressing. The more hydrophobic peptides stayed in the column for longer, while the more hydrophilic (water soluble) peptides flowed through quickly. As they exited the tube, the peptides were sent to the next instrument: the mass spectrometer.
Here, the instrument created a spray of droplets containing peptides mixed with positively charged protons. The protons stuck to the peptides, giving the peptides a positive charge. Each positively charged peptide then flew through an electric field toward a detector which recorded the mass-to-charge ratio of each peptide. These masses were then compared to a peptide database to determine the identity of each peptide present in the serum sample. While traditional proteomics methods would require sample preparation to isolate each protein before determining its identity [5], the shotgun proteomics method used here was able to identify all the proteins in the sample after just a single digestion step coupled with LCMS.
Using blood from just 17 bats, Neely and coworkers identified two potentially important peptides, one from the Rh186 protein of a herpes virus, and one from the ORF1a protein of a MERS-CoV virus. Importantly, MERS-CoV and SARS-CoV viruses are two species within the same viral genus called the betacoronaviruses [4]. This was the first study to show that viral proteins can be detected from bat blood through shotgun proteomics. Previous searches of viruses in bats have used PCR to detect viral genes within the sample, just like the PCR tests used to detect SARS-CoV-2. However, PCR tests are highly specific, meaning that a virus can only be detected if scientists are specifically searching for it. With shotgun proteomics, all viral proteins located within a sample can be identified all at once, as long as that protein is known and recorded in a database. Additionally, if a protein is identified in the future, datasets from the present could be used to backtrack and follow the virus’s path, potentially from bat to livestock to human.
Although this initial study was small, sampling only 17 bats, its success at detecting viral proteins from bat blood samples demonstrates the future promise of shotgun proteomics to find and track viruses in wild animals. Hopefully, research using this technique and others will help scientists amass enough data to better understand where zoonotic diseases come from and how they transmit to humans to prepare and protect against future infectious diseases.
[1] https://www.cdc.gov/onehealth/basics/zoonotic-diseases.html
[2] https://www.nature.com/articles/d41586-021-00865-8
[3] https://www.healthline.com/health/trypsin-function

