Article: Evolving artificial metalloenzymes via random mutagenesis
Authors: Hao Yang, Alan M. Swartz, Hyun June Park, Poonam Srivastava, Ken Ellis-Guardiola, David M. Upp, Gihoon Lee, Ketaki Belsare, Yifan Gu, Chen Zhang, Raymond E. Moellering and Jared C. Lewis
Journal: Nature Chemistry
For most scientists, changing things at random is not the best approach to solving a problem. However, when dealing with complex biological systems, it can sometimes lead to phenomenal results. It may be hard to believe that random changes can actually generate a desirable outcome, yet this is how life has evolved its complexity over billions of years. One wonder of the evolution of life is protein folding. Imagine the challenge of trying to connect hundreds of paperclips together into a chain to create a specific functional shape, such as a sphere or rope. When dealing with proteins, the building blocks are amino acids instead of paperclips, and the shapes that are made become the machines and structures that maintain all life on our planet. The authors of the paper discussed in this Chembite attempt to mimic life by evolving artificial (non-natural) proteins using biological randomization techniques.
Background on Proteins
Proteins are combinations of hundreds, sometimes thousands, of smaller molecules known as amino acids that bond together in a process known as polymerization. There are roughly 20 of these amino acids that occur naturally. By switching out one amino acid for another, scientists can change the properties of these biological polymers.
It’s simple to change a single amino acid – that’s only 20 variations if you want to test every amino acid. It gets complicated once you begin selecting more than 1 position within the protein to alter. With only two positions to change, there are already 400 different combinations (20^2). Since proteins are typically composed of hundreds and even thousands of amino acids, this task quickly becomes daunting (20^100 is an unimaginably large number, and don’t even try 20^1000). Therefore, randomization is sometimes necessary to probe greater depths within the protein because it gives researchers the ability to easily change multiple parts of the protein. This is especially useful when information about the protein is scant. Exact structural information is difficult to obtain and is not available for every known protein, in particular for artificial proteins like the enzymes discussed here (such as how they fold together into a useful shape and the areas in the enzyme where the metal catalyst and reactants may bind). There is interest in these types of artificial proteins because the creation and evolution of artificial enzymes could enable completely new types of reactions that are useful for both chemists and biologists.
Artificial Metalloenzymes
In this research, the proteins investigated are artificial metalloenzymes (ArMs), which are composed of a protein scaffold that can tightly bind to synthetic rarer metal cofactors, such as ruthenium, rhodium, iridium, among others (Figure 1). This metal cofactor can catalyze new types of reactions, since these rarer metals are typically not found in natural enzymes and exhibit different activities from the naturally-occurring metal cofactors. Specifically, these authors are concerned with the selectivity of these new catalysts. Selectivity involves how the protein holds the reactants in a certain position so that the catalyst can only react with a specific version (or enantiomer) of the chemical of interest. One way to envision this process is to think about putting a glove onto one of your hands: the left-handed glove has the snuggest fit on your left hand, so it’s far more favorable for you to put it on your left hand than your right. Similarly, only the “left-handed” version of a molecule will fit into the active site of the enzyme, leading to more reactions with that type of molecule.
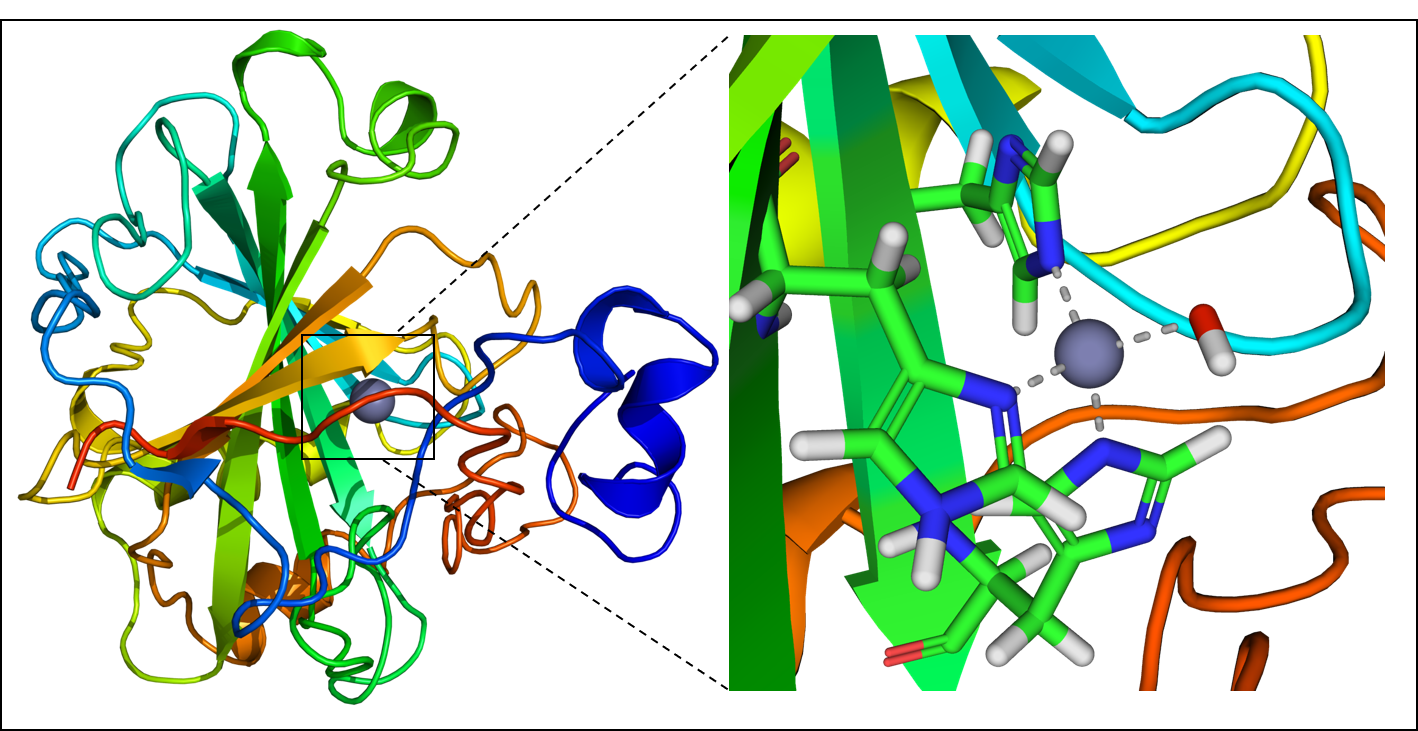
Introducing Random Changes
To generate random changes (mutations) in the protein sequence, the authors must manipulate the DNA sequence that codes for the enzyme. They do this by using proteins that do a very bad job of copying DNA. Naturally-occurring proteins known as polymerases replicate DNA and in most organisms, the process has very little error. However, scientists have both discovered and created polymerases that copy DNA very poorly, leaving mistakes in the sequence. In this case, the mistakes are useful, because they assist in the directed evolution process by generating random mutations throughout the DNA and randomizing certain portions of the protein.
The process of protein randomization is usually iterative: the researchers will go through round-by-round, isolating the best variants from each round. These best variants will then be used in subsequent rounds, undergoing further randomization. The key to finding a useful protein from the randomization process is the screening method. For enzymes, scientists generally test for the chemical that the enzyme makes (the product). In this case, the authors use di-rhodium complexes as their synthetic metal cofactor. These di-rhodium complexes are capable of catalyzing the cyclopropanation reaction shown in Figure 2. By detecting the product from Figure 2, the researchers can discover which variations of the protein work better (Figure 3). It is interesting to note that many of the randomized amino acids are not close to the active site in the middle of the image, suggesting that even a distant change alters the way the metal cofactor and reactant bind within the protein. There are a wide range of reactions catalyzed by these dirhodium complexes, which the authors demonstrate in the publication.
Take-Home Message
It is truly remarkable that these researchers are improving human-designed proteins through artificial evolution. By showing that the evolution of synthetic biomolecules with new metal catalysts can be directed through random changes, the authors lay the ground work for further evolution of artificial biocatalysts that may be capable of catalyzing entirely new types of reactions.
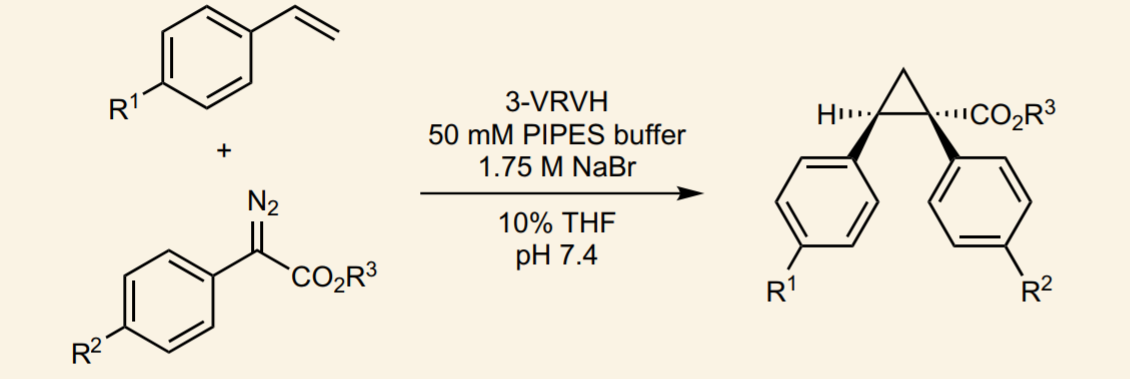
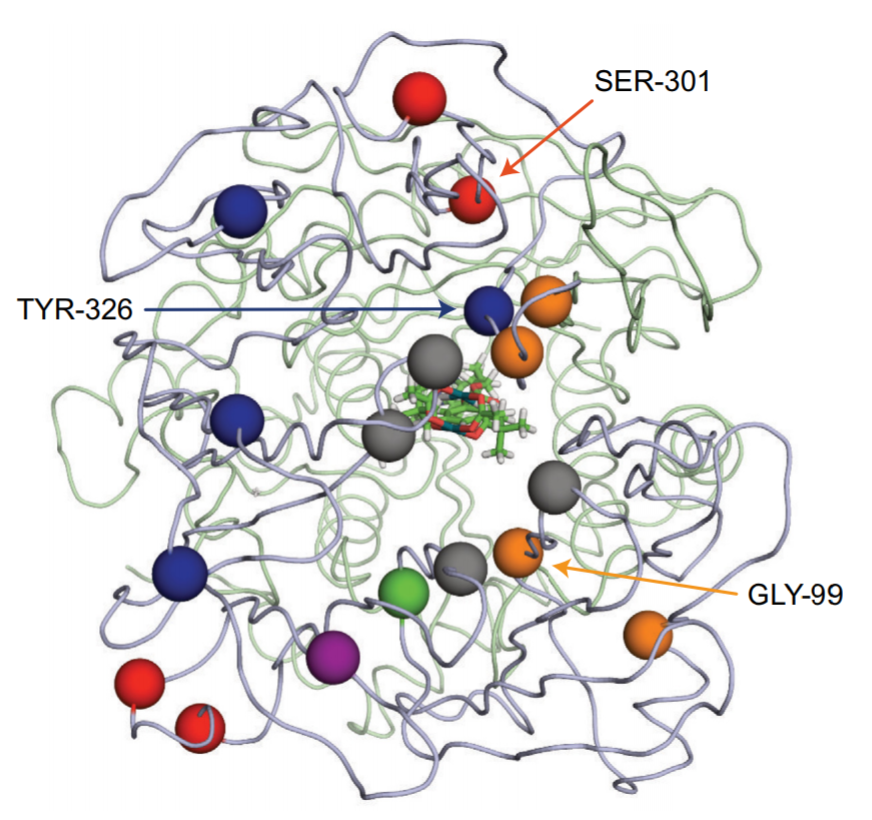
Image Credits
Featured image: Giovanni Pintori, Olivetti Historical Archive.

