RNA is an important biomolecule present in our body that participates in various biological activities. For example, mRNA acts as messenger of genetic information playing vital role during disease progression and cell development. tRNA and rRNA actively participate to synthesize proteins, which are another important biomolecule present in our body. Tracking the RNAs to understand their working mechanism is one of the mainstays of recent scientific advancement as this can provide intricate details about cellular functions and diseases. Although several detection techniques (e.g. Northern blotting, polymerase chain reaction) exist, these techniques generally require several steps and cannot detect RNAs in vivo. However, it is important to visualize the RNAs in live cells noninvasively to fully understand their functions.
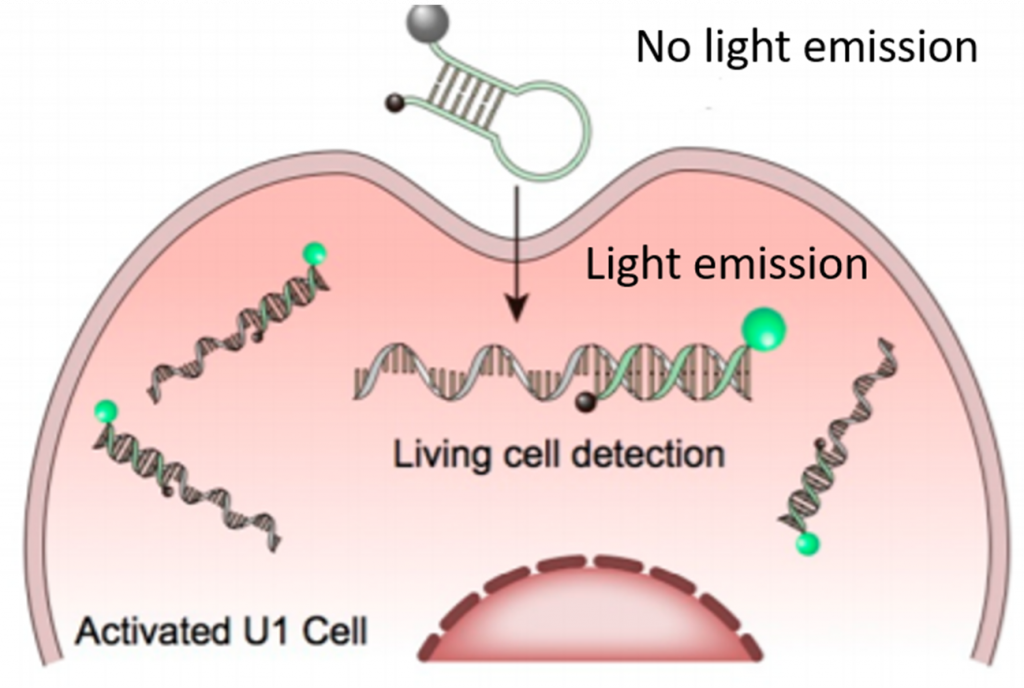
To address this, scientists can attach light-emitting molecules to RNAs to monitor these molecules inside cells. Usually, quantum dots (QDs) are used as they are small in size and easily modifiable thus does not disrupt the cellular environment. In this work, a group of researchers from China wanted to exploit this QD-based strategy to image a single RNA molecule. However, they would not be able to attach QDs to the RNAs being produced inside a living cell in real time, so they instead attached a QD to DNA: a molecule that would selectively bind the RNA they wanted to study. This DNA-QD conjugate would include a DNA sequence complementary to the RNA they wanted to study, as well as an attached “quenching” molecule that would prohibit the QD from emitting light. When the DNA-QD conjugate comes into contact with its complementary RNA, the QD is released from the quencher. The released QD then tags along with the RNA of interest while emitting light (Scheme 1). In this way, it acts as a sensor material to detect RNA.
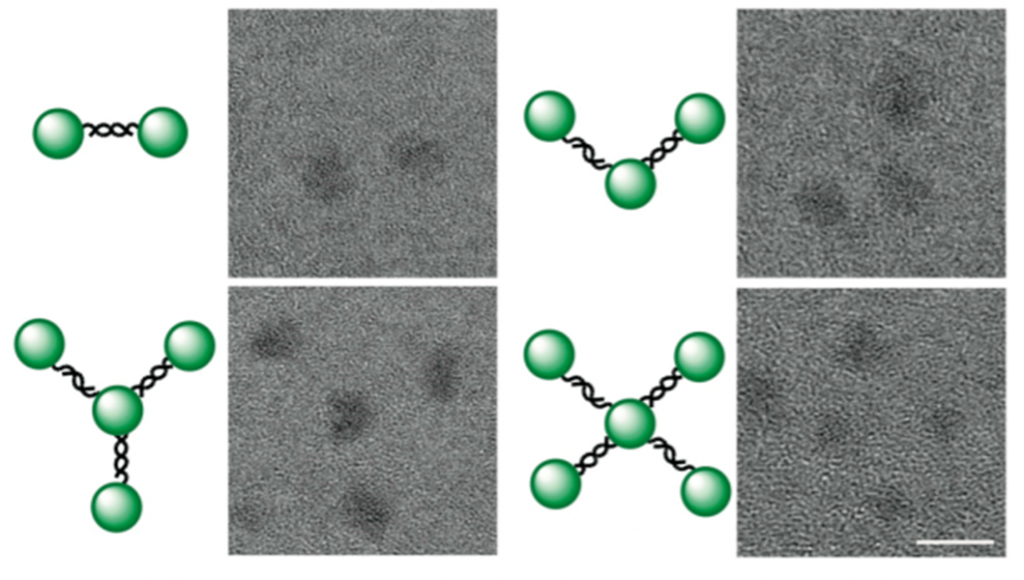
To see which DNA-QD conjugate would work best as an RNA probe, scientists want to make different kinds of conjugates, sometimes with multiple dots or DNA strands. In this work, different types of DNA-QD conjugates were prepared by varying DNA concentration and structural modification in the preparation step. Formation of different structures was then monitored using Transmission Electron Microscopy (Figure 1). Dynamic Light Scattering, which provides information about size and charge of the particles in solution, showed formation of larger particles with more negative charges when DNA was conjugated to the QDs. As DNA is intrinsically negatively charged, these experiments proved attachment of DNA with the QDs.
Initial studies were performed using spectroscopic techniques to explore the feasibility of the DNA-QD conjugates for live imaging. The QDs showed significant fluorescence intensity initially, but after addition of DNA the fluorescence intensity decreases when DNA-QD conjugates form due to the presence of quencher. Interaction of DNA-QD conjugates with RNA, again gives back the fluorescence intensity. These materials were found to be more stable than other similar reagents against cleaving enzymes, indicating they could be used within live cells.
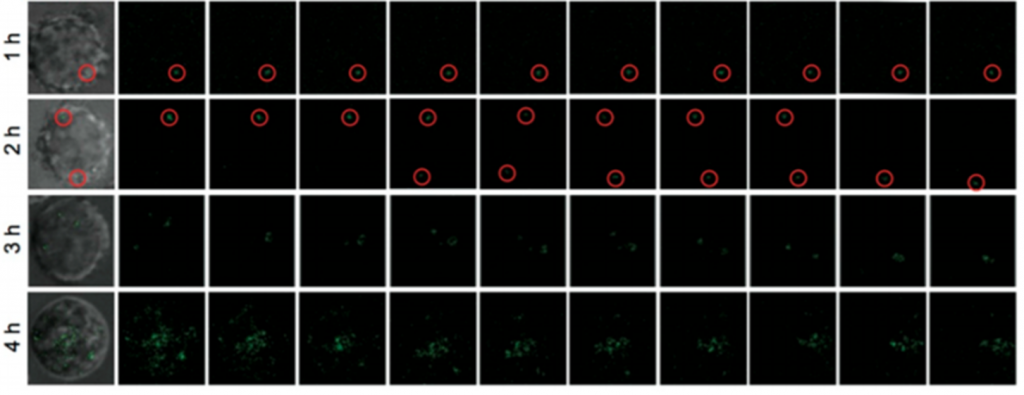
Live imaging experiments were then performed against HIV-1 integrated cells. The DNA-QDs tested emitted light with higher efficiency and longer time as compared to related materials. Initially, in the first 30 minutes, just a little fluorescence was detected, and it started to increase with time. After around 4h, most of the cell was brightened up as expected (Figure 2). Due to gradual slow growth of fluorescence, within the first 15 minutes the scientists could effectively detect a single RNA. This ability to detect a single molecule could be extremely useful in tracking one RNA molecule’s path within a cell.
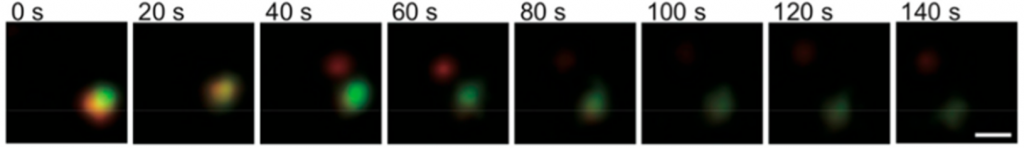
To fully understand their potential applications, the scientists then monitored a dynamic process within a single HIV-1 virus. Unlike most organisms from bacteria to humans, HIV-1 and some other viruses carry their genetic information as RNA, not DNA. Therefore, this virus is the perfect system for the new RNA probe. In the experiment, DNA-QD conjugates were made to interact with RNA during the HIV-1 Assembly processes , forming an HIV-1-QD conjugate. To deliver these HIV-1-QD conjugates within the cell, a protective protein shell was necessary which is generally called capsid. These capsids were functionalized with another color marker (red colored dye named ReAsH) to monitor the uncoating of HIV-1-QD conjugates within the cell from their protein shell. Now the whole material ‘HIV-1-QD-ReAsH’ has two colored sections: QD containing HIV-1 (green color) and ReAsH containing capsid (red color) (Figure 3). When these conjugates were inserted within a live cell, HIV-1 part was uncoated, triggering the release of green colored QD conjugates from red colored capsid that were monitored using microscopy. In this way, live tracking of the dynamic processes of single virus was achieved.
With increasing number of genomic treatments of disease and other important biological applications, it is essential to develop imaging tools to visualize genetic material, including both DNA and RNA molecules. This new RNA probing technique can provide a novel platform to image nucleic acids within live cells, following the RNAs involved in human health and disease.
Reference
- Ma, Y.; Mao, G.; Huang, W.; Wu, G.; Yin, W.; Ji, X.; Deng, Z.; Cai, Z.; Zhang, X.; He, Z.; Cui, Z. Quantum Dot Nanobeacons for Single RNA Labeling and Imaging. J. Am. Chem. Soc. 2019, 141, 13454-13458.
Feature image was modified from (image by Kevin Payravi, reused under ‘Creative commons Attribution-Share Alike 3.0 Unported’ license) https://commons.wikimedia.org/wiki/File:Microscope_icon_(black).svg

