| Title | Identification and characterization of the key enzyme in the biosynthesis of the neurotoxin β-ODAP in grass pea |
| Author | Moshe Goldsmith, Shiri Barad, Maor Knafo, Alon Savidor, Shifra Ben-Dor, Alexander Brandis, Tevie Melhman, Yoav Peleg, Shira Albeck, Orly Dym, Efrat Ben-Zeec, Ranjit S. Barbole, Asaph Aharoni, Ziv Reich |
| Journal | Journal of Biological Chemistry |
| Year | 2022 |
| URL | https://www.jbc.org/article/S0021-9258(22)00246-0/fulltext |
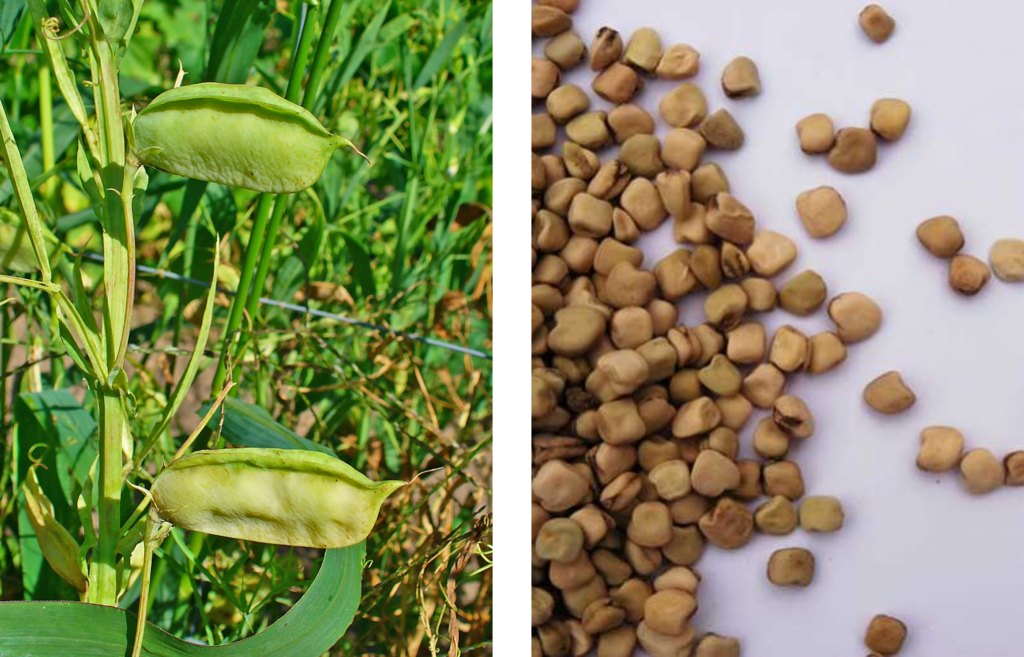
Though many are familiar with the neurotoxins in castor beans (ricin) and hemlock (coniine), you may not have heard of β-N-oxalyl-L-α, β-diaminopropionic acid, abbr. β-ODAP. β-ODAP is an unusual compound that overexcites neurons, traveling all the way from the ingestion point to the point by easily crisscrossing the blood-brain barrier. The neurotoxin originates from the grass pea (Fig 1), a foraged and cultivated legume in South Asia and sub-Saharan Africa. During drought or famine, the hardy legume is commonly ground into flour or used as animal fodder. The crop staple, known also as khersari dal, was originally banned in India 60 years ago because of the lower limb paralysis that it caused, with long-time consumption leading to irreversible neurolathyrism. Now with the ban lifted in 2016, plant bioengineers are increasingly interested in cultivating a strain of grass pea that does not produce the toxin to guarantee hardy foodstuffs for times of drought.

Scientists first figured out how the chemical β-ODAP functions in 1964. Chemists at the Indian Institute of Science extracted the toxic compound from seeds of the pea plant, known to be enriched in β-ODAP, and characterized its properties. Based on the chemical structure, bioengineers modified strains of the plant to express an enzyme that could degrade a precursor to β-ODAP, leading to transgenic plants with significantly less neurotoxin. However, to date, there is no bioengineered pea plant that is completely neurotoxin-free – mainly because the biosynthetic process of the chemical was unknown. That is, until a recent 2022 paper published by Israeli scientists Goldsmith et al. revealed the enzyme responsible for it, which they named β-ODAP synthase (BOS).
How did they do it? By tricking the enzyme into revealing itself.
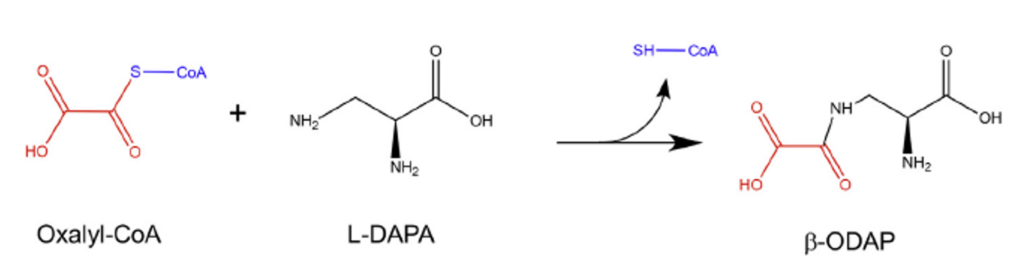
Scientists had previously figured out that the final step of the β-ODAP biochemical pathway was the fusion of two chemicals, oxalyl-CoA and L-DAPA (Figure 2). The difficulty in finding the bonafide BOS was that the particular reaction to form β-ODAP is not very specific. Multiple enzyme types can facilitate the reaction, such as ligases (enzymes that make a bond between two compounds), transferases (transfer of one functional group to another), or even CoA binding enzymes (because of CoA involvement). The scientists first had to narrow down their options for what could be the BOS by extracting as many proteins as possible from β-ODAP enriched parts of the plant, like the seeds and sprouts.
The scientists crushed seeds into a paste and used high speed centrifugation to separate out lipids, nucleic acids, and carbohydrates from the targeted proteins. They separated batches of the proteinaceous mixture into individual fractions by their hydrophobic or water-repellant properties. Afterwards, they performed an initial screening for β-ODAP activity by incubating the fractions with oxalyl-CoA and L-DAPA, looking for the disappearance of L-DAPA. As L-DAPA is consumed by the enzyme and transformed to β-ODAP, there is less “free”, color-absorbing L-DAPA in solution. The scientists would then take the batches that had the most activity and separate them based on another intrinsic property again, until only single-protein-pure fractions with high β-ODAP activity were left.
This screening process eventually resulted in 219 (!) protein candidates, which were identified using peptide mass fingerprinting. However, the Israeli scientists narrowed down the list further by including only those that were predicted by their sequence to be in the relevant enzyme classes and posses a CoA binding site. This resulted in 18 candidates, more manageable for their in-depth, downstream analyses. The corresponding genes for each of the 18 proteins were copied into bacterial cells and used to make pure protein. Of the 18 final candidates, only one possessed substantial β-ODAP activity. After characterizing the protein structure and using molecular simulation to investigate the enzymatic mechanism, the enzyme was officially named the elusive β-ODAP synthase or BOS of the grass pea.
Goldsmith et al.’s work completes our knowledge as to the biochemical production of the grass pea’s neurotoxin β-ODAP. Over ₤1.2 million was awarded to the United Kingdom’s John Innes Foundation for their work in 2018, which was based on a low-neurotoxin-producing grass pea strain. In identifying the specific β-ODAP synthase protein and its corresponding gene, this opens the potential for an entirely neurotoxin-free strain. Cultivating this bioengineered pea plant would greatly expand the world’s ability to combat hunger in times of famine and drought without losing nutritional value. No longer does the grass pea’s BOS elude us – instead, we are the new boss of BOS.

