Article Title: Use of a scaffold peptide in the biosynthesis of amino acid-derived natural products
Authors: Chi P. Ting, Michael A. Funk, Steve L. Halaby, Zhengan Zhang, Tamir Gonen, Wilfred A. van der Donk
Journal: Science
Year: 2019
DOI: 10.1126/science.aau6232
Although we often don’t think about it, we owe much of our lifestyle to microbes. These tiny bugs are chemical workhorses responsible for giving cheese its distinct taste and churning out antibiotics. As such, scientists are interested in studying microbial chemistry and discovering novel natural products. This typically involves identifying an interesting trait (such as antibiotic activity) and then finding the enzymes responsible for making this compound. With the advent of DNA sequencing technologies, however, we can now do this in the reverse by first identifying interesting stretches of DNA (called biosynthetic gene clusters) and then figuring out what compounds are created, or biosynthesized.
One particularly interesting class of natural products is ribosomally synthesized and posttranslationally modified peptides (RiPPs). Nearly all of these compounds begin as a precursor peptide that is subsequently modified by other enzymes encoded in the gene cluster. The final products (entirely modified peptides) have diverse uses, including as antimicrobials, food preservatives, and feed additives. .
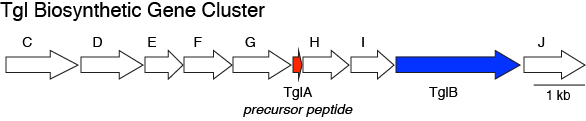
Ting and coworkers identified a peculiar RiPP-encoding gene cluster, the Tgl biosynthetic gene cluster (Fig 1). They were able to show that the precursor peptide (TigA) is used catalytically to generate an amino acid-derived natural product (3-thiaglutamate) (Fig 2). Unlike the typical RiPP system where the entire precursor peptide is converted into the final natural product, TigA is regenerated after producing 3-thiaglutamate. Excitingly, the authors were also able to show that this paradigm extends beyond this one case. By identifying additional biosynthetic gene clusters with similar peculiar motifs, they similarly showed that the ammosamides are biosynthesized using similar logic.
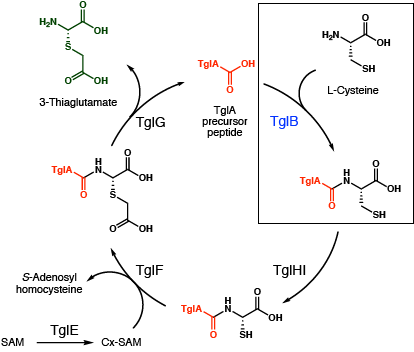
The researchers first came across the Tgl gene cluster after noticing an odd tailoring enzyme called TglB. Unlike similar proteins, TglB was shorter and missing sections needed to perform its usual chemistry of adding and eliminating the amino acid L-glutamate. Protein mass spectrometry later confirmed that instead of modifying an amino acid within the precursor peptide, TglB added the amino acid L-cysteine to the end of TglA (Fig 2, boxed).

Using similar methods, Ting and coworkers were able to piece together the roles of the other enzymes in the Tgl gene cluster (TglEFGHI) and show that this terminal cysteine is ultimately converted into the compound 3-thiaglutamate. The key experiment was identifying the stereochemistry of the final product. This was accomplished via cryo-electron microscopy (cryo-EM) method microcrystal electron diffraction (MicroED), a technique developed in 2013. Small crystal samples are bombarded with x-rays, resulting in a diffraction pattern (Fig 3) that can be computationally converted into a chemical structure. The biological role of the final product, 3-thiaglutamate, is not yet known. The authors’ hypothesis is that this microbial product mimics host plant signaling molecules and blocks the plant’s response to invading pathogens.

Altogether, this report showcases the utility of studying microbial chemistry by going from gene cluster to function. This reverse methodology enables scientists to prioritize studying new functions that have not been identified before. Through this work, the researchers discovered that a precursor peptide can be used to produce microbial metabolites catalytically. Instead of wasting cell resources constructing a large precursor peptide to produce just one compound, microbes have evolved an efficient recycling method. This key discovery stemmed from wanting to understand a peculiar enzyme of unknown function.