Article Title: A fasciclin protein is essential for laccase-mediated selective phenol coupling in sporandol biosynthesis.
Authors: Thiele, W.; Obermaier, S.; Müller, M.
Journal: ACS Chem. Biol.
Year: 2020
DOI: 10.1021/acschembio.0c00025
Fungi are some of nature’s most prolific chemists, capable of synthesizing a variety of complex molecules. These secondary metabolites, or natural products, often have useful biological activities, including as antimicrobials, viral inhibitors, psychedelics, and chemotherapies. As such, many research groups are focused on discovering new natural products and understanding how fungi create, or biosynthesize, these compounds. By doing so, others can take advantage of this newfound knowledge and use it to improve production yields or engineer more potent molecules.
In this new work, Thiele and coworkers uncover the process by which the fungus Chrysosporium merdarium synthesizes an anti-parasitic chemical called sporandol. Intriguingly, this compound contains fused biaryls (see blue areas in Figure 1A). Prior work had shown that this reaction is usually catalyzed by either a laccase, cytochrome P450 enzyme, or peroxidase (which are all different classes of enzymes). In the case of sporandol, however, a different mechanism is at play. Through a combination of microbiology and biochemistry, the authors show that a new type of enzyme is responsible. More specifically, both a laccase and a fasciclin domain-containing (fas) protein of unknown function are required for this characteristic transformation.
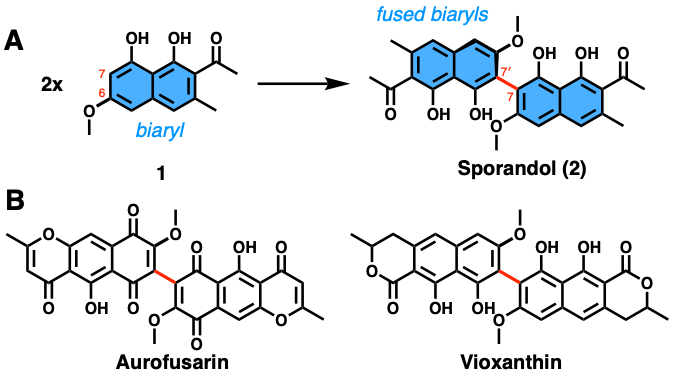
The first step was to identify the genes that are responsible for sporandol production. The authors sequenced the whole genome of C. merdarium, the fungus that produces sporandol. Guided by the structure of sporandol, the authors were able to make an educated guess about what types of enzymes were needed. This enabled them to identify the Spo biosynthetic gene cluster, a stretch of DNA of clustered genes (Figure 2A). Most notable in the cluster were two genes: SpoL, which encodes a laccase, and SpoF, which encodes a fas protein.
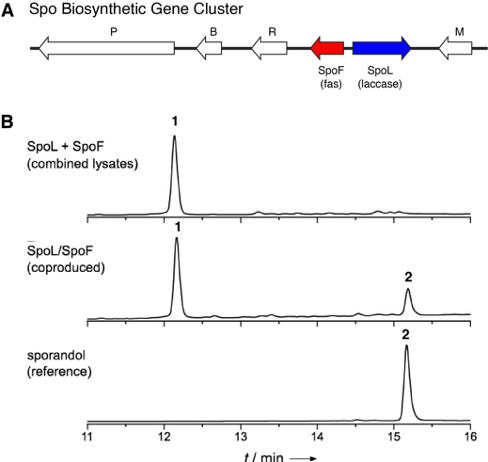
The authors transferred these genes to two different organisms (Aspergillus niger and Escherichia coli) and were able to show that a combination of the enzymes SpoP and SpoM produced compound 1 (the direct precursor of sporandol). This was verified by comparing the experimentally made compound with a known synthetic standard. The most interesting question, however, was how the dimerization of 1 to form 2 (the desired sporandol) occurs. To study this, the authors expressed the enzymes individually in A. niger. Incubating the enzymes alone with compound 1 did not lead to any product formation. This was also the case when two enzymes were simply mixed together. However, when the two enzymes were co-expressed together, 2 could be detected (Figure 2B). The authors speculate that both enzymes may be needed for proper folding of the enzymes or that a protein complex was being formed.
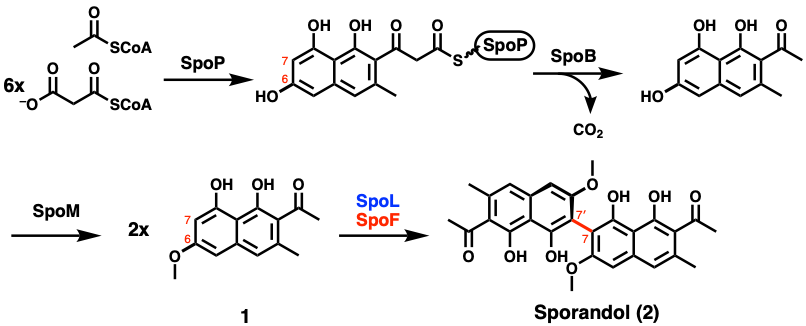
In the end, a new biosynthetic hypothesis for how sporandol is produced was developed (Figure 3). Through this work, the authors showed that nature has a variety of methods to install similar biaryl architectures. This is advantageous, as it expands the chemical toolbox of enzymes that scientists can use to design their own biosynthetic pathways. Continued efforts in this line of research will enable further discovery of novel natural products and potentially therapeutics.

